
Identification of Haplotypes
Associated With Resistance to
Bacterial Cold Water Disease in
Rainbow Trout Using Whole-Genome
Resequencing
Sixin Liu
1
*, Kyle E. Martin
2
, Guangtu Gao
1
, Roseanna Long
1
, Jason P. Evenhuis
1
,
Timothy D. Leeds
1
, Gregory D. Wiens
1
and Yniv Palti
1
1
National Center for Cool and Cold Water Aquaculture, Agricultural Research Service, United States Department of Agriculture,
Kearneysville, WV, United States,
2
Troutlodge Inc., Sumner, WA, United States
Bacterial cold water disease (BCWD) is an important disease in rainbow trout aquaculture.
Previously, we have identified and validated two major QTL (quantitative trait loci) for BCWD
resistance, located on chromosomes Omy08 and Omy25, in the odd-year Troutlodge May
spawning population. We also demonstrated that marker-assisted selection (MAS) for
BCWD resistance using the favorable haplotypes associated with the two major QTL is
feasible. However, each favorable haplotype spans a large genomic region of 1.3–1.6 Mb.
Recombination events within the haplotype regions will result in new haplotypes associated
with BCWD resistance, which will reduce the accuracy of MAS for BCWD resistance over
time. The objectives of this study were 1) to identify additional SNPs (single nucleotide
polymorphisms) associated with BCWD resistance using whole-genome sequencing
(WGS); 2) to validate the SNPs associated with BCWD resistance using family-based
association mapping; 3) to refine the haplotypes associated with BCWD resistance; and 4) to
evaluate MAS for BCWD resistance using the refined QTL haplotypes. Four consecutive
generations of the Troutlodge May spawning population were evaluated for BCWD
resistance. Parents and offspring were sequenced as individuals and in pools based on
their BCWD phenotypes. Over 12 million SNPs were identified by mapping the sequences
from the individuals and pools to the reference genome. SNPs with significantly different
allele frequencies between the two BCWD phenotype groups were selected to develop SNP
assays for family-based association mapping in three consecutive generations of the
Troutlodge May spawning population. Among the 78 SNPs derived from WGS, 77
SNPs were associated with BCWD resistance in at least one of the three consecutive
generations. The additional SNPs associated with BCWD resistance allowed us to reduce
the physical sizes of haplotypes associated with BCWD resistance to less than 0.5 Mb. We
also demonstrated that the refined QTL haplotypes can be used for MAS in the Troutlodge
May spawning population. Therefore, the SNPs and haplotypes reported in this study
provide additional resources for improvement of BCWD resistance in rainbow trout.
Keywords: rainbow trout, bacterial cold water disease, haplotype, SNP, MAS, QTL, whole genome
sequencing (WGS)
Edited by:
Nguyen Hong Nguyen,
University of the Sunshine Coast,
Australia
Reviewed by:
Antti Kause,
Natural Resources Institute Finland
(Luke), Finland
Yulin Jin,
Emory University, United States
*Correspondence:
Sixin Liu
Specialty section:
This article was submitted to
Livestock Genomics,
a section of the journal
Frontiers in Genetics
Received: 05 May 2022
Accepted: 06 June 2022
Published: 23 June 2022
Citation:
Liu S, Martin KE, Gao G, Long R,
Evenhuis JP, Leeds TD, Wiens GD and
Palti Y (2022) Identification of
Haplotypes Associated With
Resistance to Bacterial Cold Water
Disease in Rainbow Trout Using
Whole-Genome Resequencing.
Front. Genet. 13:936806.
doi: 10.3389/fgene.2022.936806
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368061
ORIGINAL RESEARCH
published: 23 June 2022
doi: 10.3389/fgene.2022.936806

INTRODUCTION
The global demand for seafood has roughly doubled since the
start of the 21
st
century, and will likely double again by 2050
(Naylor et al., 2021). Rainbow trout (Oncorhynchus mykiss) is one
of the most widely cultured cold freshwater fish, with production
on every continent except Antarctica. The global production of
rainbow trout was about 917,000 tons in 2019 (FAO, 2021).
Outbreaks of infectious disease are one of the major challenges for
rainbow trout production and welfare. Bacterial cold water
disease (BCWD), caused by Flavobacterium psychrophilum,is
a frequent disease in rainbow trout (Nematollahi et al., 2003;
Starliper, 2011; Loch and Faisal, 2015). Commercial vaccines for
BCWD are not available yet. Use of licensed antibiotics for
BCWD treatment increases production cost and antibiotic
resistant pathogens may emerge.
Use of genetic resistance is an effective approach to control
BCWD in rainbow trout. A rainbow trout line with improved
resistance to BCWD has been developed by using family-based
selection (Leeds et al., 2010; Wiens et al., 2013a; Wiens et al.,
2018). Recently, multiple studies have demonstrated that
genomic selection (GS) can substantially improve the accuracy
of selection for BCWD resistance in rainbow trout. Vallejo et al.
(2017a) reported that GS using a 57K SNP (single nucleotide
polymorphism) genotyping array (Palti et al., 2015a) can double
the accuracy of selection for BCWD resistance in a commercial
breeding population. To reduce the cost of genotyping, the
accuracy of GS for BCWD resistance was evaluated with low-
density SNP panels. The accuracy of GS remained substantially
higher than pedigree-based selection when using 70 SNPs
associated with QTLs (quantitative trait locus) for BCWD
resistance (Vallejo et al., 2018). To reduce the cost of BCWD
phenotyping, it has recently been reported that the accuracy of GS
for BCWD resistance without model retraining in the subsequent
generation remained higher than pedigree-based selection
(Vallejo et al., 2021).
To fully exploit the genetic resistance to BCWD, extensive
genetic mapping studies were conducted to identify and validate
QTLs for BCWD resistance in rainbow trout. Fraslin et al. (2018)
used both immersion and intramuscular injection methods to
evaluate double haploids derived from a cross between two
rainbow trout isogenic lines, and 15 QTLs for BCWD
resistance were identified. Also, two QTLs for BCWD
resistance were identified after a natural disease outbreak on a
French farm (Fraslin et al., 2019). At the USDA National Center
for Cool and Cold Water Aquaculture, we initially used full-sib
mapping families to identify and validate QTL for BCWD
resistance (Wiens et al., 2013b; Vallejo et al., 2014a; Vallejo
et al., 2014b; Palti et al., 2015b; Liu et al., 2015). With the
advancement of genomic resources available in rainbow trout
such as a SNP genotyping array (Palti et al., 2015a) and a
reference genome (Pearse et al., 2019), we performed a
genome-wide association study (GWAS) to detect QTL for
BCWD resistance (Vallejo et al., 2017b) in the 2013
generation of the Troutlodge May spawning population. Three
QTL for BCWD resistance with moderate-large effects, located on
chromosomes Omy03, Omy08, and Omy25, were identified. In a
follow-up study the three QTLs were validated in the 2015
generation of the Troutlodge May spawning population (Liu
et al., 2018). In the same study it was shown that SNP
haplotypes associated with the two major QTL on
chromosomes Omy08 and Omy25 can be used for marker-
assisted selection (MAS) for BCWD resistance. However, the
two favorable haplotypes for the two major QTL on
chromosomes Omy08 and Omy25 span regions of 1.3 and
1.6 Mb, respectively. Recombination events within the
haplotype regions may result in new haplotypes associated
with BCWD resistance, which will reduce the accuracy of
MAS for BCWD resistance over time. Thus, it is important to
identify and validate additional SNPs associated with the two
major QTL for BCWD resistance with a goal to reduce the
physical size of haplotypes associated with BCWD resistance.
Whole-genome sequencing (WGS) is a powerful tool to
discover SNPs and to identify causative genes for traits of
interest. With the recent rapid reduction in the cost of next
generation sequencing, the use of WGS has become more
common in genetic studies of salmonids (Gao et al., 2018;
Narum et al., 2018; Thompson et al., 2020; Liu et al., 2021
).
The objectives of this study were 1) to identify additional SNPs
associated with BCWD resistance using WGS; 2) to validate the
SNPs associated with BCWD resistance using family-based
association mapping; 3) to refine the haplotypes associated
with BCWD resistance; and 4) to evaluate MAS for BCWD
resistance using the refined QTL haplotypes.
MATERIALS AND METHODS
Five Consecutive Ge nerations of the
Troutlodge May Spawning Strain
Troutlodge, Inc., has four rainbow trout strains (Liu et al., 2017)
named by their peak spawning months. All samples used in this
study were from the May spawning strain. Five consecutive
generations (Table 1) were used in this study. The samples
used for WGS were from the 2013 and 2015 generations.
Selected families from three consecutive generations, 2015,
2017, and 2019, were used for association analyses of BCWD
resistance. Fish of the 2021 generation were used to evaluate MAS
for BCWD resistance. Previously, each Troutlodge strain had two
populations, odd-year and even-year populations. The odd-year
and even-year May spawning populations have been merged into
one population since the 2019 generation.
BCWD Challenge Experiments
BCWD challenge experiments were conducted in four
consecutive generations, 2015, 2017, 2019, and 2021
(Supplementary Table S1). Fish (80–99 days post-hatch) were
challenged by intraperitoneal injection of Flavobacterium
psychrophilum strain CSF259-93 using the established protocol
described in detail by Hadidi et al. (2008). Mortalities were
collected daily for 21 days after intraperitoneal injection. Both
survival days (DAYS), the number of days to death after BCWD
challenge, and survival status (STATUS), 2 for dead fish and 1 for
survivors at day 21, were recorded for each fish. Each family of the
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368062
Liu et al. Haplotypes Associated With BCWD Resistance

2015 and 2017 generations was evaluated for BCWD resistance
using two replicate tanks (3 L tank with a water flow rate of 1 L
per minute) with 40 fish per tank, and the details have been
reported in our previous publications (Vallejo et al., 2017a; Liu
et al., 2018). For the 2019 generation, we increased the number of
fish challenged per family to 3 or 4 replicate tanks (40 fish per
tank) based on fish availability. The 2021 families were pooled
and challenged as described below to evaluate MAS for BCWD
resistance.
Sequencing of 40 Parents of the 2015 and
2017 Generations
Based on the BCWD survival rates of the 2015 and 2017 families
and parental haplotypes for the two targeted QTL regions,
Omy08 and Omy25, we selected 40 (Supplementary Table
S2) parents for individual sequencing and pooled sequencing.
First, we sorted the families within each generation by BCWD
survival rate from high to low. The parents of top 20 families were
assigned to a BCWD resistant (R) group, and the parents of
bottom 20 families were assigned to a BCWD susceptible (S)
group. Then, the QTL haplotypes of these parents were
reconstructed for the two QTL regions using the same SNP
sets and method reported in Liu et al. (2018). The parents of
the 2015 and 2017 generations were selected to target for the
Omy08 and Omy25 BCWD QTL, respectively. For each
generation, we selected 10 R parents that are fixed for the
favorable haplotype for the targeted QTL and have at least one
favorable haplotype for the other QTL. We also selected 10 S
parents without any favorable haplotype for the targeted QTL.
Thus, a total of 40 parents were selected for WGS with a targeted
genome coverage of 15x per sample. In addition to sequencing of
individuals, we also pooled equal amount of DNA per fish by
BCWD groups within each generation for pooled sequencing.
The targeted genome coverage per pool was 30x. The raw
sequences of the parents were deposited in sequence read
archive (SRA) under BioProject PRJNA681179.
Sequencing of Pooled Offspring of the 2015
Generation
The sequencing of parents described above might be biased
because both BCWD survival rates and QTL haplotypes were
used to select the samples used for sequencing. Thus, we decided
to use BCWD phenotype as the only criteria to select samples for
additional pooled sequencing. Among the 138 families of the
2015 generation evaluated for BCWD resistance, we selected 60
families with intermediate BCWD survival rates that ranged from
24% to 71% for sequencing. For each of the 60 families, we
selected the first four fish that died after day 3 (to avoid fish died
from injection injury or stress) and four random survivors. Each
of the four dead fish or survivors was randomly assigned to one of
the four corresponding BCWD status pools. In total, we had four
DNA pools of dead fish (S pools) and four DNA pools of
survivors (R pools). Equal amount of DNA per sample was
pooled for sequencing with a targeted genome coverage of 45x
per DNA pool. The sequences of the pooled offspring were
deposited under BioProject PRJNA830380.
Whole-Genome Sequencing and SNP
Identification
DNA was extracted from fin clips following the manufacturer’s
recommended protocols for AutoGenprep 965 (Autogen,
Holliston, MA, United States). Whole-genome DNA
sequencing libraries were prepared using the KAPA
HyperPrep kit (KAPA Biosystems, Wilmington MA), and
were sequenced in paired-end (2 × 150 bp) mode on an
Illumina HiSeq X sequencer. The sequence reads were mapped
to rainbow trout reference genome GCF_002163495.1 (Pearse
et al., 2019) using BWA-MEM algorithm (Li, 2013), and
alignments were converted to BAM (Binary sequence
Alignment/Map) format using SAMtools v1.11 (Li et al.,
2009). PCR duplicates were marked and removed using Picard
v2.18.2 (http://broadinstitute.github.io/picard/). Following our
previously published SNP calling and filtering pipeline (Gao
et al., 2018), SNPs were called using program freebayes v1.3.1
(Garrison and Marth, 2012), and were annotated using program
SnpEff v4.3 (Cingolani et al., 2012).
Identification of SNPs in the QTL Regions
For the sequence data of parents for the 2015 and 2017
generations, we used a sliding window approach to identify
SNPs in the BCWD QTL regions. We used a window size
10,000 bp and a step size 5,000 bp to calculate the fi xation
index (Fst) value for each window. For individual sequencing,
program VCFtools v0.1.16 (Danecek et al., 2011) was used to
calculate Fst value for each sliding window. For pooled
sequencing, program PoPoolation2 (Kofler et al., 2011) was
used to calculate Fst value for each sliding window. Only
TABLE 1 | Summary of samples from five consecutive generations of the Troutlodge May spawning population used in this study.
Generation Study Comment
2013 WGS Selected parents for the 2015 generation were used for WGS (Individuals and pools; N = 20)
2015 WGS, SNP validation 1) Selected parents (Individuals and pools; N = 20) and families (pools; N = 480) were used for WGS; 2) Validate SNPs
associated with BCWD resistance using 60 families (10 offspring per family; N = 600) with intermediate BCWD survival rates
2017 SNP validation Validate SNPs associated with BCWD resistance using 60 families (10 offspring per family; N = 600) with intermediate
BCWD survival rates
2019 SNP validation Validate SNPs associated with BCWD resistance using 9 families (92 offspring per family; N = 828) with intermediate BCWD
survival rates
2021 MAS Evaluation of MAS using Pooled BCWD challenge (50 families; 20 fish per family; N = 1,000)
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368063
Liu et al. Haplotypes Associated With BCWD Resistance

windows with at least 15 SNPs were included to identify windows
with significantly different allele frequencies (empirical p <
0.0001) between the R and S groups. For individual
sequencing, program VCFtools was also used to calculate Fst
value for each SNP with MAF (minor allele frequency) greater
than 0.05.
To identify SNPs with significantly different allele frequencies
between the pools of R and S offspring from the 2015 generation,
Fisher’s exact tests were performed using the program
PoPoolation2 with the following settings: -min-coverage 40,
--max-coverage 400 and--min-count 10. To correct for multiple
tests, SNPs with p-values less than 4.05 × 10
–9
(Bonferroni-
correction for 12,338,978 SNPs) were considered as significant.
SNP Genotyping
Among the SNPs identified by WGS, we selected a set of SNPs in
the Omy08 and Omy25 QTL regions for association analyses. The
SNPs were selected for assay design because they met one or more
of the following criteria: 1) The Fst values between R and S
parents were high; 2) The p-values of Fisher’s test for different
allele frequencies between R and S pools of the 2015 generation
were low; 3) The SNPs had high or moderate effects based on SNP
annotation; 4) SNPs are near the six SNPs used in our previous
haplotype analysis (Liu et al., 2018). The sequences of the selected
SNPs were submitted to Fluidigm (South San Francisco, CA) for
assay design. After a preliminary evaluation of assay quality using
a subset of mapping samples of the 2015 generation, we
assembled a panel of 96 SNPs (Supplementary Table S3)to
genotype the mapping samples of three consecutive generations,
2015, 2017, and 2019.
We followed the SNP genotyping protocol described in our
previous study (Liu et al., 2016). Briefly, DNA samples were pre-
amplified, and the pre-amplified products were diluted and used
for genotyping with 96.96 Dynamic Array IFCs (Integrated
Fluidic Circuits). The arrays were read using EP1 system, and
genotypes were called automatically using Fluidigm SNP
genotyping analysis software 4.1 with a confidence threshold
of 85. The genotype clusters were examined by eye for each assay,
and any wrong calls or no calls were corrected manually. The
computer program PedCheck (O’Connell and Weeks, 1998) was
used to identify genotypes with Mendelian inheritance errors
between parents and offspring. Seven SNPs (Supplementary
Table S3) were removed from association analysis due to poor
genotype clusters or high rates of genotype discrepancies between
parents and offspring.
Family-Based Association Mapping of
BCWD Resistance
The program PLINK 1.9 (Chang et al., 2015) was used for family-
based association mapping to validate SNPs associated with BCWD
resistance (p < 0.01). The procedure QFAM was used to analyze the
phenotypic data DAYS, and the PERM option was used to correct
the family structure. The procedure TDT (transmission
disequilibrium test) was used to analyze the binary phenotype
STATUS. The association analyses were performed for each of
the three consecutive generations, 2015, 2017, and 2019.
Haplotype Association Analysis of BCWD
Resistance
We used three criteria to select three SNPs per QTL region for
haplotype association analysis. 1) The selected SNPs are highly
associated with BCWD resistance based on single SNP
association analysis; 2) The MAF for each selected SNP is
greater than 0.2; and 3) The three SNPs for each QTL region
span a genomic region less than 0.5 Mb according to r ainbow
trout reference genome GCF_002163495.1 (Pearse e t al.,
2019). Based these thr ee criteria, three SNPs , P489 , P194,
and P355, were selected for the Omy08 QTL, a nd three
SNPs, P420, P430, and P212, w ere selected for the Omy25
QTL. The same families used for single SNP analysis described
above were also used for haplotype association analysis. The
haplotypes for each fish were constructed using Beagle 5.1
(Browning and Browning, 2007), and haplotypes with
frequency larger than 0.1 were retained for haplotype
association analysis. Program PLINK1.9 was used for
haplotype association analysis following the sam e method
for family-based association analysis of single SNP as
described above.
Evaluation of MAS for BCWD Resistance in
the 2021 Generation
The parents for the 2021 generation were genotyped with 96
SNPs (Supplementary Table S3), and haplotypes for the
Omy08 and Omy25 QTL regions were constructed using
thesameSNPsandmethoddescribedabove.The163
families of the 2021 generation were sorted by the total
count of favorable haplotypes from high to low. The top 25
families were assigned to the high haplotype group, and the
bottom 25 families were assigned to low haplotype group. We
pooled 10 fish per family by haplotype grou ps, and the 2 50 fish
were challenged with BCWD in a 40 L tank with a water flow
rate of 2.5 L per minute. There were two replicate tanks for
each haplotype group. So, a total of 500 fish were challenged
per haplotype group. To test the BCWD survival difference
between the high and low favorable haplotype groups, a log-
rank test was performed using the survival package ( Therneau,
2021) a vailable in R version 4.1.2 (R Core Team, 2021).
RESULTS
Identification of Genomic Regions
Associated With BCWD Resistance
Using WGS
The 20 parents for the 2015 generation used for sequencing were
selected to target the Omy08 QTL for BCWD resistance. For
individual sequencing, all windows with significantly different Fst
between R and S parents were in a region between 73.2 and
78.2 Mb on chromosome Omy08 (Figure 1A). For pooled
sequencing, except two windows on chromosome Omy05 and
one window on chromosome Omy13, all the other windows with
significantly different Fst between R and S pools were in a region
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368064
Liu et al. Haplotypes Associated With BCWD Resistance

between 73.2 and 78.0 Mb on chromosome Omy08 (Figure 1B).
Thus, we selected SNPs in the region from 73.2 to 78.2 Mb to
validate SNPs associated with the BCWD QTL on chromosome
Omy08.
The 20 parents for the 2017 generation used for sequencing
were selected to target the Omy25 QTL for BCWD resistance. For
individual sequencing, all windows with significantly different Fst
between R and S parents were in a region between positions 16.5
and 40.1 Mb on chromosome Omy25 (Figure 2A). For pooled
sequencing, except one window on chromosome Omy05 and one
window on chromosome Omy12, all the other windows with
significantly different Fst between R and S pools were located on
chromosome Omy25 in a region between 18.9 and 41.0 Mb
(Figure 2B). Thus, we selected SNPs in the region from 16.5
to 41.0 Mb to validate SNPs associated with the BCWD QTL on
chromosome Omy25.
FIGURE 1 | Distribution of Fst between R and S groups of parents for the 2015 generation of the Troutlodge May spawning population. (A) sequencing of
individuals; (B) sequencing of pooled samples. The red horizontal lines represent the threshold for significantly different Fst (empirical p < 0.0001).
FIGURE 2 | Distribution of Fst between R and S groups of parents for the 2017 generation of the Troutlodge May spawning population. (A) sequencing of
individuals; (B) sequencing of pooled samples. The red horizontal lines represent the threshold for significantly different Fst (empirical p < 0.0001).
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368065
Liu et al. Haplotypes Associated With BCWD Resistance

To avoid the potential bias of samples used for sequencing just
described above, we also sequenced pooled offspring of the 2015
generation. The samples were selected with BCWD phenotypes alone,
and they were pooled by survival status. After mapping the sequence
reads to the reference genome, 12.3 million SNPs were identified.
Based on Fisher’s test for each SNP, 21 SNPs with significantly
different allele frequencies between the R and S pools were
identified (Figure 3), and they were located on chromosomes
Omy04, Omy05, Omy08, Omy11, Omy16, Omy17, Omy20, and
Omy25. Only chromosomes Omy08 and Omy25 had more than three
significant SNPs, and these significant SNPs were located in similar
QTL regions to those identified by WGS of parents as described above.
Validation of SNPs Associated With BCWD
Resistance
Among the 89 SNPs used for association analysis in three consecutive
generations, 2015, 2017, and 2019, 85 SNPs were associated with
BCWD resistance in at least one of the three generations (Table 2).
Also,77outofthe85validatedSNPswereidentified via WGS
reported in this study. Among the 4 SNPs, P161, P176, P316, and
P490, that were not associated with BCWD resistance in this study,
P490 was the only SNP derived from WGS.
Refined Haplotypes Associated With BCWD
Resistance
The 85 SNPs associated with BCWD resistance validated above
allowed us to reduce the physical size of the haplotypes associated
with BCWD resistance. Using the three criteria described in the
method section, we selected SNPs, P489, P194, and P355, for the
Omy08 QTL, and SNPs, P420, P430, and P212, for the Omy25
QTL. The three selected SNPs span a region less than 0.5 Mb.
Thus, we reduced the sizes of haplotypes by about two-thirds. The
results of haplotype association analysis are summarized in
Table 3. For the Omy08 QTL, haplotype CGG was associated
with BCWD resistance in all three generations, which increases
the number of survival days and reduces the risk of death from
BCWD. Similarly, haplotype GGG on chromosome Omy25 was
also associated with BCWD resistance in all three generations,
which increases the number of survival days and reduces the risk
of death from BCWD. The other two haplotypes (Table 3) were
associated with BCWD susceptibility. Because the goal of selective
breeding is to improve BCWD resistance, we will focus on the two
haplotypes associated with BCWD resistance and refer to them as
favorable haplotypes for BCWD resistance.
Evaluation of MAS for BCWD Resistance
The 25 families from the 2021 generation with high or low counts
of favorable haplotypes for the two major QTL regions had an
average of 6.48 and 2.56 favorable haplotypes per family,
respectively. The BCWD survival rates for the pooled families
with high or low counts of favorable haplotypes were 83.6% and
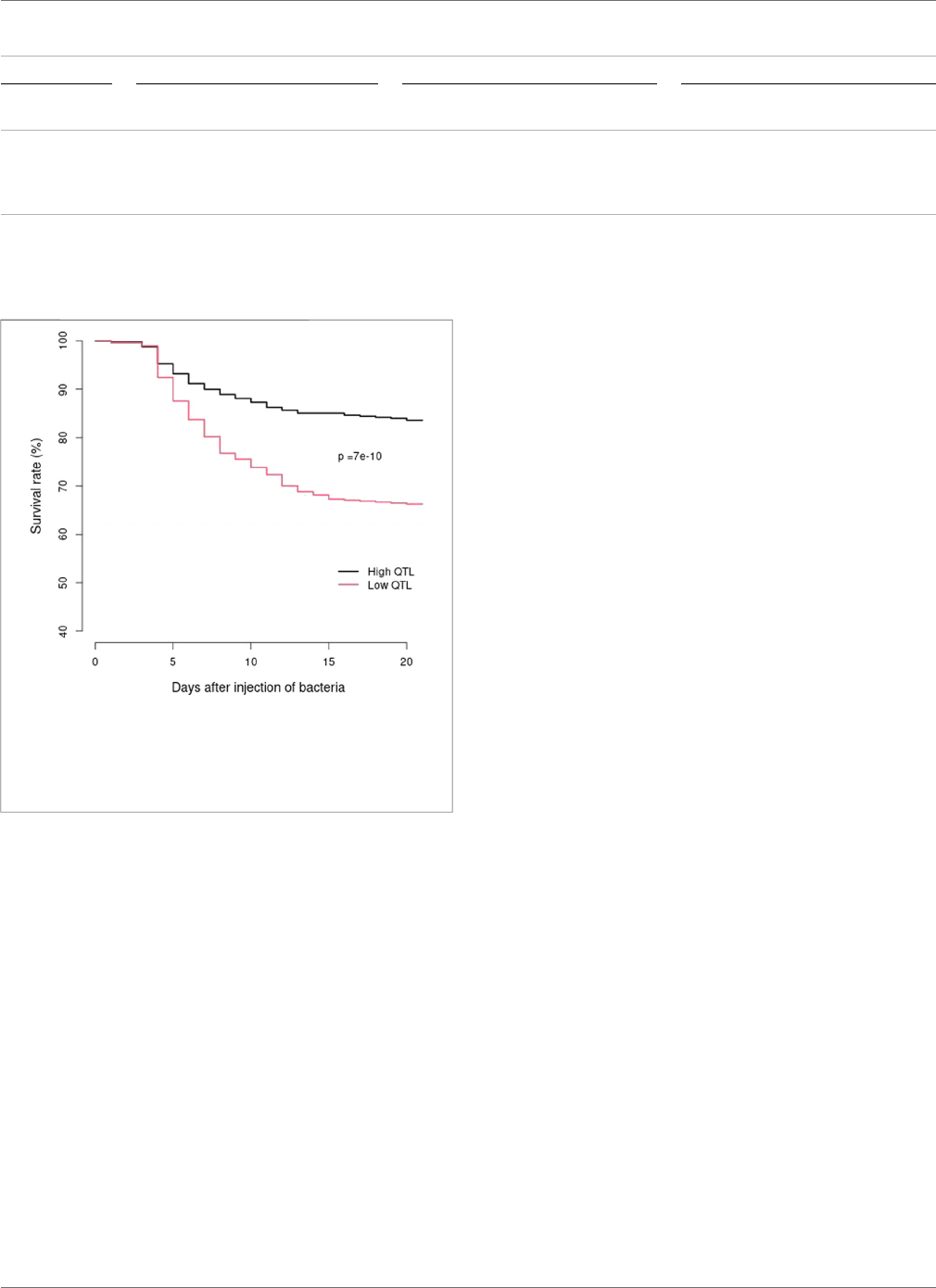
66.2%, respectively. Survival analysis demonstrated that the two
BCWD survival curves were significantly different (p = 7e-10)
(Figure 4) for the two groups of families with high or low counts of
favorable haplotypes. Thus, the favorable haplotypes can be used to
select families with improved BCWD resistance.
DISCUSSION
In this study, we used WGS to identify additional SNPs associated
with the two major QTL for BCWD resistance, and 77 SNPs
identified from WGS were validated by association mapping in
three consecutive generations of the Troutlodge May spawning
population. The additional SNPs associated with BCWD resistance
allowed us to refine the favorable haplotypes associated with
BCWD resistance. We demonstrated that the refined favorable
QTL haplotypes can be used for MAS for BCWD resistance in the
Troutlodge May spawning population.
Identification of SNPs Associated With
BCWD Resistance Using WGS
Among the 78 SNPs derived from WGS, only SNP P490 was not
associated with BCWD resistance in this study. The high SNP
validation rate was largely due to several factors. 1) The samples
used for sequencing were selected with two methods, BCWD
phenotypes together with QTL haplotypes or BCWD phenotypes
alone; 2) We used two sequencing strategies, sequencing of
individuals and pooled samples; 3) The SNPs selected for
assay design were filtered by multiple criteria as described in
the method section; 4) We removed SNPs with poor genotype
quality or SNPs were not associated with BCWD resistance based
on a preliminary study with a sub-set of mapping samples. Due to
the genotyping platform used in this study, only 96 SNPs
including both SNPs derived from WGS and SNPs reported in
our previous study (Liu et al., 2018) were used for association
mapping. However, analysis of WGS revealed many more SNPs
FIGURE 3 | Manhattan plot of pooled offspring of the 2015 generation of
the Troutlodge May spawning population. The red horizontal line represents
the significance threshold (Bonferroni-correction for 12,338,978 SNPs).
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368066
Liu et al. Haplotypes Associated With BCWD Resistance
TABLE 2 | Validation of SNPs associated with BCWD resistance in three consecutive generations of the Troutlodge May spawning population.
Generation 2015 2017 2019
Chr. SNP
a
Source Position
b
Allele1 Allele2 Effect
c
p
(DAYS)
OR
d
p
(STATUS)
Effect p
(DAYS)
OR p
(STATUS)
Effect P (DAYS) OR P (STATUS)
3 P178 Liu et al. (2018) 57059195 G A 2.8 3.7E-04 0.6 1.1E-04 1.3 NS
e
0.7 7.1E-03 1.9 NS 0.8 NS
8 P318 WGS 75,054,110 T G 3.6 7.9E-05 0.7 1.7E-03 1.9 2.1E-03 0.7 2.7E-03 −0.3 NS 1.0 NS
8 P319 WGS 75,056,552 G C 3.5 3.2E-05 0.7 3.3E-03 1.9 4.9E-03 0.7 6.1E-03 −0.9 NS 1.0 NS
8 P323 WGS 75,064,254 A T 2.8 1.6E-03 0.7 NS 1.9 2.3E-03 0.7 8.5E-03 −0.3 NS 1.0 NS
8 P489 WGS 76,457,672 A C −4.0 1.0E-04 1.4 NS -2.9 6.3E-05 1.4 NS −2.2 NS 1.3 4.5E-03
8 P329 WGS 76,691,975 T A −4.0 3.0E-06 1.5 2.7E-04 -2.3 1.7E-03 1.3 NS −2.5 NS 1.6 1.6E-05
8 P330 WGS 76,692,073 T A −4.0 3.0E-06 1.5 2.2E-04 -2.2 2.5E-03 1.3 NS −2.6 NS 1.6 1.1E-05
8 P332 WGS 76,696,519 C A − 3.9 2.0E-06 1.4 4.8E-03 -2.9 2.4E-05 1.4 NS −2.1 NS 1.2 NS
8 P194 Liu et al. (2018) 76,747,151 A G −4.0 2.0E-06 1.6 3.5E-05 -2.3 2.0E-03 1.3 NS −2.2 NS 1.5 1.3E-04
8 P341 WGS 76,758,777 A G −4.0 2.0E-06 1.5 4.8E-04 −2.6 2.4E-04 1.4 5.0E-03 −1.2 NS 1.2 NS
8 P342 WGS 76,765,345 A T −3.9 3.0E-06 1.5 5.8E-04 −2.5 6.4E-04 1.4 7.3E-03 −2.5 8.6E-03 1.3 5.0E-03
8 P346 WGS 76,815,705 G T −3.9 3.0E-06 1.5 5.8E-04 −2.1 8.1E-04 0.9 NS −2.5 8.6E-03 1.5 4.1E-05
8 P347 WGS 76,844,443 G A −3.8 4.2E-05 1.4 6.8E-03 −2.4 1.2E-03 1.3 NS −1.8 NS 1.3 NS
8 P350 WGS 76,853,682 G T −4.1 1.2E-05 1.6 3.2E-04 −2.1 5.9E-03 1.2 NS −1.6 NS 1.6 3.0E-05
8 P354 WGS 76,874,914 C T −3.8 4.2E-05 1.4 6.8E-03 −2.4 1.3E-03 1.3 NS −2.2 NS 1.5 5.2E-05
8 P355 WGS 76,875,468 A G −3.8 4.2E-05 1.4 6.8E-03 −2.3 1.3E-03 1.5 4.7E-04 −2.2 NS 1.5 5.2E-05
8 P356 WGS 76,879,188 A G −3.8 4.2E-05 1.4 6.8E-03 −2.4 1.3E-03 1.3 NS −2.2 NS 1.5 5.2E-05
8 P357 WGS 76,879,806 C A − 3.8 4.2E-05 1.4 6.8E-03
−2.4 1.3E-03 1.3 NS −2.2 NS 1.5 5.4E-05
8 P191 Liu et al. (2018) 78,064,599 A G −3.1 3.2E-04 1.2 NS −2.8 2.3E-03 1.3 NS −1.2 NS 1.3 NS
13 P494 WGS 42,160,393 C A -4.1 2.0E-04 1.4 7.5E-03 −2.5 6.4E-04 1.4 7.3E-03 −2.4 NS 1.4 7.1E-03
25 P381 WGS 17,246,875 C A −6.2 2.0E-06 1.6 NS −2.5 9.1E-03 1.6 2.4E-03 −3.0 8.6E-03 1.5 4.2E-05
25 P382 WGS 17,272,388 A G −5.7 3.0E-06 1.5 NS −2.6 1.7E-03 1.6 7.5E-04 −2.8 8.6E-03 1.5 2.4E-05
25 P383 WGS 17,893,051 T C 2.5 NS 0.7 NS −0.8 NS 1.2 NS 1.4 NS 0.8 8.3E-03
25 P384 WGS 17,944,754 T A 3.7 4.2E-05 0.6 5.2E-03 2.5 5.4E-03 0.6 7.2E-03 4.5 NS 0.3 1.2E-06
25 P386 WGS 18,159,829 A G 3.7 4.2E-05 0.6 4.1E-03 2.1 9.5E-03 0.7 NS 4.5 NS 0.3 1.2E-06
25 P230 Liu et al. (2018) 18,192,885 G T 2.5 NS 0.7 NS 2.6 5.1E-04 0.6 2.1E-04 2.7 NS 0.6 2.7E-06
25 P391 WGS 19,063,776 A G −5.5 3.0E-06 2.1 8.0E-06 −3.0 1.4E-04 1.7 3.1E-04 −3.0 NS 1.6 1.7E-06
25 P392 WGS 19,131,282 A G −6.3 1.0E-06 2.0 5.4E-08 −3.7 3.0E-06 1.7 3.5E-05 −2.6 NS 1.5 1.3E-06
25 P393 WGS 19,285,578 T C −6.7 1.0E-06 1.9 1.7E-05 −3.4 1.5E-04 1.6 1.2E-03 −2.6 NS 1.4 2.1E-04
25 P395 WGS 19,356,106 A T −5.5 1.0E-06 1.9 2.8E-05 −2.9 1.9E-04 1.6 3.1E-04 −3.4 2.3E-03 1.7 2.8E-09
25 P398 WGS 19,480,661 A G 3.9 3.0E-05 0.6 1.4E-03 2.8 1.5E-03 0.6 7.2E-03 4.8 NS 0.3 4.2E-07
25 P399 WGS 19,525,929 G T -6.3 1.0E-06 2.1 5.9E-08 −3.8 1.0E-06 1.7 2.0E-05 −2.7 NS 1.5 1.1E-06
25 P214 Liu et al. (2018) 19,553,268 C A 4.1 1.0E-06 0.7 1.1E-03 2.8 2.0E-04 0.7 6.9E-03 3.2 3.8E-03 0.6 1.4E-08
25 P402 WGS 19,652,654 T A -6.1 1.0E-06 2.0 2.1E-07 -3.6 1.0E-06 1.7 2.6E-05 -3.3 2.3E-03 1.7 4.0E-09
25 P404 WGS 19,682,358 G A 4.1 1.0E-06 0.7 1.1E-03 2.9 8.1E-05 0.7 5.0E-03 3.5 7.2E-03 0.6 4.8E-09
25 P409 WGS 19,886,570 C T 3.9 1.4E-05 0.6 2.0E-04 1.2 NS 0.8 NS 1.5 NS 0.7 NS
25 P412 WGS 20059482 C A − 5.9 1.0E-06 1.9 1.1E-06 −3.6 4.0E-06 1.7 1.2E-04 −2.6 NS 1.5 5.1E-07
25 P413 WGS 20,082,809 T C −5.5 1.0E-06 1.9 5.7E-05 −3.0 1.9E-04 1.6 3.1E-04 −3.6 4.3E-03 1.8 5.9E-09
25 P415 WGS 20,153,489 C T 4.1 1.5E-05 0.6 1.6E-03 1.6 NS 0.7 7.2E-03 3.3 NS 0.5 2.1E-05
25 P417 WGS 20,228,457 C T 3.5 7.0E-06 0.7 4.1E-03 1.8 NS 0.9 NS 2.4 NS 0.7 8.6E-05
25 P420 WGS 20,306,455 G A 4.0 1.0E-06 0.7 5.8E-03 3.2 2.1E-05 0.8 NS 3.6 7.2E-03 0.5 1.0E-09
25 P495 WGS 20,307,827 C A −6.0 1.1E-05 1.9 4.4E-05 −3.7 1.0E-06 1.7 1.2E-04 −2.5 NS 1.5 2.2E-06
25 P424 WGS 20,403,696 C T − 5.9 1.0E-06 1.9 1.1E-06
−3.6 1.0E-06 1.7 1.2E-04 −2.5 NS 1.5 4.9E-07
25 P425 WGS 20,437,940 C T − 6.2 2.9E-05 1.6 NS −2.8 3.0E-03 1.6 4.6E-03 −3.2 NS 1.6 8.9E-05
25 P426 WGS 20,465,120 G A −5.7 2.9E-05 1.6 NS −2.6 NS 1.4 NS −2.5 NS 0.9 NS
25 P429 WGS 20,570,095 C T − 6.6 8.5E-06 1.8 4.3E-03 −3.4 NS 1.3 NS 0.0 NS 1.1 NS
(Continued on following page)
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368067
Liu et al. Haplotypes Associated With BCWD Resistance

TABLE 2 | (Continued) Validation of SNPs associated with BCWD resistance in three consecutive generations of the Troutlodge May spawning population.
Generation 2015 2017 2019
Chr. SNP
a
Source Position
b
Allele1 Allele2 Effect
c
p
(DAYS)
OR
d
p
(STATUS)
Effect p
(DAYS)
OR p
(STATUS)
Effect P (DAYS) OR P (STATUS)
25 P430 WGS 20,591,515 T G − 5.1 9.4E-05 1.9 1.6E-04 −2.9 2.3E-04 1.6 1.4E-03 −3.3 NS 1.8 1.2E-07
25 P431 WGS 20,647,628 A G −5.1 9.4E-05 1.9 1.6E-04 −2.9 2.3E-04 1.6 1.4E-03 −3.4 NS 1.8 4.6E-08
25 P433 WGS 20,722,720 A G −5.9 1.0E-06 1.9 1.1E-06 −3.6 1.0E-06 1.7 1.2E-04 −2.6 NS 1.5 1.5E-06
25 P212 Liu et al. (2018) 20,751,780 T G −5.9 1.0E-06 1.9 1.6E-06 −3.6 1.0E-06 1.6 1.5E-04 −2.6 NS 1.5 1.5E-06
25 P435 WGS 20,770,014 T G − 6.0 1.0E-06 1.9 1.1E-06 −3.7 1.0E-06 1.7 1.2E-04 −2.6 NS 1.5 1.4E-06
25 P436 WGS 20,797,381 C A −4.0 1.3E-04 1.6 1.1E-04 −3.1 7.0E-06 1.4 9.2E-04 −2.1 NS 1.4 2.2E-05
25 P437 WGS 20,814,089 G A −5.6 1.0E-06 1.8 1.3E-04 −3.1 8.1E-05 1.7 1.8E-04 −3.5 4.3E-03 1.7 3.3E-09
25 P438 WGS 20,905,605 T C −6.1 1.0E-06 1.8 3.1E-06 −3.8 1.0E-06 1.8 3.2E-07 −3.2 NS 1.7 1.5E-09
25 P439 WGS 20,918,179 C T − 5.1 9.4E-05 1.9 1.6E-04 −2.9 2.0E-04 1.6 1.4E-03 −3.2 NS 1.7 4.4E-08
25 P440 WGS 20,937,844 A T 3.9 3.0E-05 0.6 2.4E-03 2.5 5.9E-03 0.7 NS 4.8 NS 0.3 4.2E-07
25 P443 WGS 21,034,905 T G 4.2 1.2E-05 0.6 2.1E-03 1.4 NS 0.7 7.2E-03 4.9 NS 0.3 1.4E-07
25 P446 WGS 21,118,315 A C −5.5 3.0E-06 1.8 1.3E-04 −2.7 5.6E-04 1.5 6.0E-03 −3.5 4.3E-03 1.7 1.2E-08
25 P228 Liu et al. (2018) 21,146,360 T G −5.6 2.0E-06 1.9 8.7E-05 −3.0 8.7E-05 1.7 1.8E-04 −3.4 2.3E-03 1.7 2.1E-09
25 P447 WGS 21,164,811 A G −6.2 2.9E-05 1.6 NS −2.8 4.1E-03 1.6 5.8E-03 −3.2 NS 1.6 1.2E-04
25 P497 WGS 21,245,059 T C 3.0 2.9E-03 0.7 NS 2.7 3.1E-03 0.7 NS 4.9 NS 0.3 1.4E-07
25 P448 WGS 21,245,244 T C 3.8 7.4E-05 0.6 2.4E-03 2.7 3.1E-03 0.7 NS 2.5 NS 0.5 6.0E-04
25 P498 WGS 21,260,524 T G 3.6 3.5E-04 0.8 NS 2.9 2.0E-05 0.7 1.3E-04 3.4 4.0E-03 0.6 9.4E-09
25 P499 WGS 21,260,707 C T 3.0 3.2E-03 0.7 NS 2.6 5.0E-03 0.7 NS 4.9 NS 0.3 9.0E-08
25 P451 WGS 21,429,395 T C −6.4 6.5E-06 1.7 4.8E-03 −3.7 3.9E-03 1.5 NS −3.8 NS 1.7 2.0E-03
25 P360 WGS 21,430,497 G T 3.4 1.5E-05 0.7 6.2E-03 1.5 NS 0.9 NS 2.7 NS 0.7 1.0E-05
25 P361 WGS 21,431,434 C T 3.7 2.0E-06 0.7 9.3E-03 3.0 1.3E-04 0.8 NS 3.0 NS 0.6 2.9E-08
25 P363 WGS 21,468,711 C G 3.8 2.0E-06 0.7 NS 2.7 3.7E-04 0.9 NS 2.9 NS 0.6 1.8E-08
25 P372 WGS 21,469,214 T G 3.9 2.0E-06 0.7 8.3E-03 3.4 1.4E-05 0.7 NS 3.5 NS 0.6 2.7E-10
25 P222 Liu et al. (2018) 21,530,601 A C 2.5 NS 0.5 NS 2.8 3.4E-03 0.7 NS 4.9 NS 0.3 9.0E-08
25 P453 WGS 21,535,437 A G −6.2 2.9E-05 1.6 NS −2.9 2.9E-03 1.6 4.6E-03 −3.2 NS 1.6 6.0E-05
25 P458 WGS 21,955,099 T A 3.7 1.3E-04 0.6 2.4E-03 2.5 2.2E-03 0.7 NS 4.9 NS 0.3 4.2E-07
25 P374 WGS 22,346,077 G T 4.0 2.0E-06 0.7 6.0E-03 3.3 9.0E-06 0.8 NS 3.6 7.2E-03 0.6 4.4E-09
25 P369 WGS 22,363,310 T G 3.6 3.0E-06 0.7 9.5E-03 3.3 1.9E-05 0.8 NS 3.2 NS 0.6 4.4E-08
25 P371 WGS 22,373,578 A C 3.6 6.0E-06 0.7 6.1E-03 2.2 4.0E-03 0.8 NS 3.3 NS 0.6 5.9E-07
25 P468 WGS 22,900,574 A C 3.3 2.7E-05 0.7 NS 2.3 1.5E-03 0.9 NS 3.5 7.2E-03 0.6 1.1E-07
25 P469 WGS 22,929,112 G A −2.8 8.7E-04 1.4 4.1E-03 −1.7 NS 1.2 NS -3.6 NS 1.6 1.9E-05
25 P470 WGS 22,954,748 T G − 5.4 2.1E-05 1.5 NS −3.4 6.3E-05 1.7 1.4E-04 -3.4 4.0E-03 1.6 3.4E-07
25 P472 WGS 23,514,396 T C 3.4 5.6E-05 0.6 1.8E-04 0.8 NS 0.8 NS -1.0 NS 1.1 NS
25 P473 WGS 23,533,980 T C 4.3 1.0E-06 0.6 1.1E-05 2.3 1.8E-03 0.6 6.1E-04 2.9 NS 0.5 1.7E-05
25 P476 WGS 23,741,596 C T 3.8 6.2E-05 0.6 2.4E-03 2.6 1.1E-03 0.7 NS 4.5 NS 0.3 2.4E-06
25 P481 WGS 24,385,764 A G 3.9 2.9E-05 0.6 2.4E-03 2.7 1.1E-03 0.7 NS 4.9 NS 0.3 1.8E-06
25 P482 WGS 24,459,019 A C 4.4 1.0E-06 0.6 1.1E-05 2.3 3.0E-03 0.7 1.2E-03 2.9 NS 0.5 3.3E-05
25 P483 WGS 25,069,717 A T −4.9 6.7E-05 1.8 9.4E-04 −2.0 NS 1.3 NS -2.3 NS 1.5 NS
25 P485 WGS 25,580,318 T G 3.9 1.3E-05 0.6 1.6E-03 2.0 NS 0.8 NS 2.5 NS 0.6 8.8E-03
a
SNPs, used for haplotype analysis (Table 3) are highlighted in bold.
b
SNP, position on rainbow trout reference genome GCF_002163495.1.
c
Allele substitution effect, positive number indicates that allele 1 increases the number of survival days, and negative number indicates that allele 1 reduces the number of survival days.
d
Odds ratio, greater than one indicates that allele 1 increases the risk of death from BCWD, and less than one indicates that allele 1 reduces the risk of death from BCWD.
e
Not significant (p > 0.01).
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368068
Liu et al. Haplotypes Associated With BCWD Resistance
that were putatively associated with BCWD resistance. Thus,
WGS is a powerful tool to identify SNPs associated with BCWD
resistance in rainbow trout.
Sequencing pools of individuals (pool-seq) is cost-effective,
and has been successfully applied to a variety of studies
(Schlotterer et al., 2014). However, pool-seq also has technical
challenges and limitations. Unequal representation of DNA
samples in the pools can cause false positive signals. The same
parental DNA samples were sequenced individually and by pool-
seq in this study. Although the genomic regions with significantly
different Fst were similar for both sequencing strategies (Figures
1, 2) for the targeted QTL regions, a few additional genomic
regions also showed significantly different allele frequencies for
the pooled samples, which was likely due to unequal
representation of DNA samples in the pools. Compared to the
results of sequencing of parents, more genomic regions showed
significantly different allele frequencies between the DNA pools
of offspring in the 2015 generation (Figure 3). Most of them are
likely false positives due to the technical challenges of pool-seq. In
addition to the possibility of unequal representation of each
offspring in the pool, unreliable BCWD phenotypes could be a
major factor since it is not possible to have replicated BCWD
challenges of an individual fish. The offspring were selected for
pool-seq on basis of BCWD survival status of individual fish. On
the other hand, the family BCWD survival rates are based on a
large number of offspring, and hence are much more reliable than
the BCWD survival status of an individual fish.
Two Robust QTL for BCWD Resistance in
Rainbow Trout
QTL validation is essential for implement of MAS in breeding
programs and identification of causative genes. The two major QTL
for BCWD resistance, located on chromosomes Omy08 and Omy25,
were initially identified in the 2013 generation of Troutlodge May
spawning population (Vallejo et al., 2017b), and were validated in
the 2015 generation of the same population (Liu et al., 2018). In this
study, we used WGS to identify additional SNPs associated with
these two major QTL, and 77 additional SNPs associated with
BCWD resistance were validated in three consecutive
generations, 2015, 2017 and 2019, of the Troutlodge May
spawning population. Thus, the two major QTL for BCWD
resistance are robust in the Troutlodge May population, and it is
worthwhile to evaluate MAS for BCWD resistance and to identify
positional candidate genes underlying the QTL.
Robust MAS for BCWD Resistance in
Rainbow Trout
We reported previously that the accuracies of retrospective MAS
for BCWD resistance using favorable haplotypes associated with
the two major BCWD QTL were equal or greater than the
accuracies of family-based selection in the same generation of
odd-year Troutlodge May spawning population (Liu et al., 2018).
In this study, we reduced the physical size of the haplotypes by
about two-thirds. We then used the refined haplotypes for MAS for
BCWD resistance in the 2021 generation of the Troutlodge May
spawning population. Based on the QTL haplotypes of the parents,
two groups of families with high or low counts of favorable
haplotypes, respectively, were selected for pooled BCWD
challenge. The two groups of families had significantly different
BCWD survival curves. It is notable that the odd-year and even-
TABLE 3 | Haplotype association analysis of BCWD resistance in three consecutive generations of the Troutlodge May spawning population.
Generation 2015 2017 2019
Chr. Haplotype Effect
a
p
(DAYS)
OR
b
p
(STATUS)
Effect P (DAYS) OR P (STATUS) Effect P (DAYS) OR P (STATUS)
8 CGG 3.8 4.0E-06 0.6 2.9E-04 2.1 2.9E-03 0.7 1.5E-02 2.8 7.6E-03 0.7 9.9E-04
8 AAA −3.9 2.7E-05 1.2 NS
c
−2.8 2.3E-03 1.1 NS −1.3 NS 1.0 NS
25 GGG 4.0 1.0E-06 0.7 4.1E-03 3.0 4.6E-05 0.8 1.4E-02 3.5 8.8E-03 0.6 3.1E-09
25 ATT −5.1 7.3E-05 1.9 1.6E-04 −2.9 1.6E-04 1.6 1.9E-03 −3.2 NS 1.7 1.1E-06
a
Positive number indicates that the haplotype increases the number of survival days, and negative number indicates that the haplotype reduces the number of survival days.
b
Odds ratio, greater than one indicates that the haplotype increases the risk of death from BCWD, and less than one indicates that the haplotype reduces the risk of death from BCWD.
c
Not significant (p > 0.05).
FIGURE 4 | BCWD survival curves of the 2021 generation of the
Troutlodge May spawnin g population. The black curve represents the families
with high counts of favorable haplotype s for the two major BCWD QTL
regions. The red curve represents the families with low counts of
favorable haplotypes for the two major BCWD QTL regions.
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 9368069
Liu et al. Haplotypes Associated With BCWD Resistance

year Troutlodge May spawning populations have been combined
into one population since the 2019 generation. Together with the
results of retrospective MAS reported previously (Liu et al., 2018),
we conclude that MAS for BCWD resistance is robust in the
Troutlodge May spawning population.
Although we focused on MAS for BCWD in this study, it is
important to note that the additional SNPs associated BCWD
resistance reported in this study should also be useful to improve
the accuracy of GS for BCWD resistance. We reported previously
that the accuracy of GS for BCWD resistance using 70 SNPs
associated with BCWD resistance was similar to the accuracy of
the whole-genome 57K SNP array (Vallejo et al., 2018).
Furthermore, it has been documented that functional and
causative variants can be used to improve the accuracy of GS
(Xiang et al., 2021). Some of the SNPs reported in this study are
located within candidate genes for BCWD resistance (see below).
Candidate Genes of QTL for BCWD
Resistance in Rainbow Trout
Our long-term goal is to identify causative genes for BCWD resistan ce
in rainbow trout. Although the refined haplotypes are associated with
resistance to BCWD, they may or may not span the QTL regions.
Thus, we arbitrarily extended 0.5 Mb on each end of the refined
favorable haplotypes associated with BCWD resistance, and then
examined protein-coding genes in the corresponding regions of
rainbow trout Arlee reference gen ome (Gao et al., 2021), which
has a better genome coverage than the previous Swanson reference
genome (Pearse et al., 2019). Based on the NCBI rainbow trout gene
annotation release 101, a total of 70 annotate d protein-coding genes
were identifiedinthetwomajorQTLregions(Supplementary
Table S4).
Among the 40 annotated protein-coding genes in the Omy08 QTL
region, multiple genes are likely related to immune responses. Both
LOC110530755 and LOC110530756 encode NACHT proteins, which
are implicated in apoptosis and MHC (major histocompatibility
complex) transcription activation (Koonin and Aravind, 2000;
Laing et al., 2008), and play important roles in activation of
animal innate immune responses to pathogen infection (Jones
et al., 2016). Furthermore, NACHT proteins such as Nod like
receptors also play an important role in activation of pyroptosis
pathway in both mammals and fish (Song et al., 2022). Three other
candidate genes, LOC110530758, LOC110530759, and
LOC110530764, encode proteins likely belonging to the signaling
lymphocytic activation molecule (SLAM) family of receptors, which in
mammals are critic al elements for both innate and adaptive immune
responses (Veillette, 2006; Cannons et al., 2011). Also, these three
SLAM genes were modestly upregulated at day 5 post challenge with
Flavobacterium psychrophilum in the study reported by Marancik
et al. (2015). In addition to the putative functions, the results of
association mapping (Table 2) also indicated that these genes are
strong candidates for the Omy08 QTL. All 8 SNPs in the candidate
gene regions (Supplementary Table S4) are significantly associated
with BCWD resistance in three consecutive generations of the
Troutlodge May spawning population (Table 2). Thus, these
immune-related genes are good candidates for the Omy08 QTL
for BCWD resistance.
Among the 30 annotated genes in the Omy25 QTL region, gene
LOC100136157, which encodes invariant chain INVX, stands out as
a promising candidate gene for this QTL. For simplicity and
consistency with rainbow trout literatures, we refer this gene as
INVX from now on. Rainbow trout INVX was initially cloned and
characterized by Fujiki et al. (2003), and is a homolog of mammalian
invariant chain genes, which play important roles in antigen
presentation (Schroder, 2016). Transcript level of INVX in
rainbow trout cell line culture was significantly increased at 96
and 120 h after immune system activation with PMA (phorbol 12-
myristate 13-acetate) (Semple et al., 2019). Also, INVX protein level
was significantly reduced at 168 h after PMA stimulation (Semple
et al., 2019). In addition to the putative function of INVX, there is
also another line of evidence supporting INVX as a candidate gene
for the Omy25 QTL. SNP P446, located in the intron of gene INVX,
was significantly associated with BCWD resistance in three
consecutive generations of the Troutlodge May spawning
population (Table 2). Therefore, we will continue to evaluate this
candidate gene using other approaches in the future.
In addition to the candidate genes highlighted above, we
would like to caution that other genes could also be candidate
genes for the BCWD QTL. Although we focused on protein-
coding genes, there are also two annotated non-coding RNA
genes in the Omy08 QTL region. We should not completely rule
out other genes in the QTL regions until we can identify with high
confi
dence the causative genes underlying the two QTL for
BCWD resistance in rainbow trout.
CONCLUSION
WGS is a powerful tool to identify additional SNPs associated with
the two major QTL for BCWD resistance in rainbow trout. The
additional SNPs allowed us to reduce the ph ysic al size of hapl ot ypes
associated with BCWD resistance. We also demonstrated that the
refined favorable QTL haplotypes can be used for MAS for BCWD
resistance in the Troutlodge May spawning population. Thus, the
additional SNPs and refinedhaplotypesassociatedwithBCWD
resistance reported in this study are useful for improvement of
BCWD resistance and for eventual identification of genes for BCWD
resistance in rainbow trout.
DATA AVAILABILITY STATEMENT
The datasets presented in this study can be found in online
repositories. The names of the repository/repositories and
accession number(s) can be found in the article/Supplementary
Material.
ETHICS STATEMENT
The animal study was reviewed and approved by Institutional
Animal Care and Use Committee, National Center for Cool and
Cold Water Aquaculture, Agriculture Research Service,
United States Department of Agriculture.
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 93680610
Liu et al. Haplotypes Associated With BCWD Resistance

AUTHOR CONTRIBUTIONS
SL and YP conceived and planned the study; KM participated in
the study planning and provided pedigree records, germplasm for
disease challenges and tissue or DNA samples from the parents;
YP, JE, TL, GW and SL coordinated, supervised and performed
the disease challenge experiments; GG mapped the sequences to
reference genome and called SNPs; SL designed genotyping plans
and RL performed the genotyping experiments; SL analyzed the
data and drafted the manuscript. All authors read and approved
the final manuscript.
FUNDING
This study was supported by Agricultural Research Service CRIS
projects 8082-32000-007 and 8082-31000-013.
ACKNOWLEDGMENTS
The authors would like to thank Kristy Shewbridge , Rya n
Lipscomb, Clayton Birkett, Jenea McGowan, Josh Kretzer,
Travis Moreland, Keira Osbourn, V anessa Panaway, and Joe
Beach for fish rearing and help with the disease challenge
experiments. Mention of trade n ames or com mercial
products in this pub lication is solely for the purpose of
providing specific informat ion and does not imply
recommendation or endorsement by the U.S. Department
of Agriculture (USDA). USDA is an equal opportunity
provider and employer.
SUPPLEMENTARY MATERIAL
The Supplementary Material for this article can be found online at:
https://www.frontiersin.org/articles/10.3389/fgene.2022.936806/
full#supplementary-material
Supplementary Table S1 | Detailed information of BCWD challenge experiments of
four consecutive generations of the Troutlodge May spawning population.
Supplementary Table S2 | The 40 parents of the 2015 and 2017 generations of the
Troutlodge May spawning population selected for sequencing.
Supplementary Table S3 | The primer sequences of 96 SNPs used for association
mapping in three consecutive generations of the Troutlodge May spawning
population.
Supplementary Table S4 | Protein-coding genes in the two major BCWD QTL
regions based on NCBI rainbow trout gene annotation release 101.
REFERENCES
Browning, S. R., and Browning, B. L. (2007). Rapid and Accurate Haplotype
Phasing and Missing-Data Inference for Whole-Genome Association Studies
by Use of Localized Haplotype Clustering. Am. J. Hum. Genet. 81, 1084–1097.
doi:10.1086/521987
Cannons, J. L., Tangye, S. G., and Schwartzberg, P. L. (2011). SLAM Family
Receptors and SAP Adaptors in Immunity. Annu. Rev. Immunol. 29 (29),
665–705. doi:10.1146/annurev-immunol-030409-101302
Chang, C. C., Chow, C. C., Tellier, L. C., Vattikuti, S., Purcell, S. M., and Lee, J. J.
(2015). Second-Generation PLINK: Rising to the Challenge of Larger and Richer
Datasets. Gigascience 4, 7. doi:10.1186/s13742-015-0047-8
Cingolani, P., Platts, A., Wang, L. L., Coon, M., Nguyen, T., Wang, L., et al. (2012).
A Program for An notating and Predicting the Effects of Single Nucleotide
Polymorphisms, SnpEff: SNPs in the Genome of Drosophila M Strain W1118;
Iso-2; Iso-3. Fly 6, 80–92. doi:10.4161/fly.19695
Danecek, P., Auton, A., Abecasis, G., Albers, C. A., Banks, E., Depristo, M. A., et al.
(2011). The Variant Call Format and VCFtools. Bioinformatics 27, 2156–2158.
doi:10.1093/bioinformatics/btr330
FAO (2021). Fishery and Aquaculture Statistics. Global Aquaculture Production
1950-2019 (FishstatJ). FAO Fisheries Division. Rome. [Online]. Available at:
http://www.fao.org/fishery/statistics/software/fishstatj/en (Accessed September
17, 2021).
Fraslin, C., Dechamp, N., Bernard, M., Krieg, F., Hervet, C., Guyomard, R., et al.
(2018). Quantitative Trait Loci for Resistance to Flavobacterium Psychrophilum
in Rainbow Trout: Effect of the Mode of Infection and Evidence of Epistatic
Interactions. Genet. Sel. Evol. 50, 60. doi:10.1186/s12711-018-0431-9
Fraslin, C., Brard-Fudulea, S., D’ambrosio, J., Bestin, A., Charles, M., Haffray, P.,
et al. (2019). Rainbow Trout Resistance to Bacterial Cold Water Disease: Two
New Quantitative Trait Loci Identified after a Natural Disease Outbreak on a
French Farm. Anim. Genet. 50, 293–297. doi:10.1111/age.12777
Fujiki, K., Smith, C. M., Liu, L., Sundick, R. S., and Dixon, B. (2003). Alternate Forms
of MHC Class II-Associated Invariant Chain Are Not Produced by Alternative
Splicing in Rainbow Trout (Oncorhynchus Mykiss) but Are Encoded by Separate
Genes. Dev. Comp. Immunol. 27, 377–391. doi:10.1016/s0145-305x(02)00119-2
Gao, G., Nome, T., Pearse, D. E., Moen, T., Naish, K. A., Thorgaard, G. H., et al.
(2018). A New Single Nucleotide Polymorphism Database for Rainbow Trout
Generated through Whole Genome Resequencing. Front. Genet. 9, 147. doi:10.
3389/fgene.2018.00147
Gao, G. T., Magadan, S., Waldbieser, G. C., Youngblood, R. C., Wheeler, P. A.,
Scheffler, B. E., et al. (2021). A Long Reads-Based De-Novo Assembly of the
Genome of the Arlee Homozygous Line Reveals Chromosomal Rearrangements
in Rainbow Trout. G3-Genes Genomes Genet. 11 (4), jkab052. doi:10.1093/
g3journal/jkab052
Garrison, E., and Marth, G. (2012). Haplotype-based Variant Detection from
Short-Read Sequencing. arXiv:1207.3907 [q-bio.GN].
Hadidi, S., Glenney, G. W., Welch, T. J., Silverstein, J. T., and Wiens, G. D. (2008).
Spleen Size Predicts Resistance of Rainbow Trout to Flavobacterium
Psychrophilum Challenge. J. Immunol. 180, 4156–4165. doi:10.4049/jimmunol.
180.6.4156
Jones, J. D., Vance, R. E., and Dangl, J. L. (2016). Intracellular Innate Immune
Surveillance Devices in Plants and Animals. Science 354 (6316), aaf6395. doi:10.
1126/science.aaf6395
Kofler, R., Pandey, R. V., and Schlotterer, C. (2011). PoPoolation2: Identifying
Differentiation between Populations Using Sequencing of Pooled DNA
Samples (Pool-Seq). Bioinformatics 27, 3435
–3436. doi:10.1093/bioinformatics/
btr589
Koonin, E. V., and Aravind, L. (2000). The NACHT Family - A New Group of
Predicted NTPases Implicated in Apoptosis and MHC Transcription Activation.
Trends Biochem. Sci. 25, 223–224. doi:10.1016/s0968-0004(00)01577-2
Laing, K. J., Purcell, M. K., Winton, J. R., and Hansen, J. D. (2008). A Genomic View
of the NOD-Like Receptor Family in Teleost Fish: Identification of a Novel NLR
Subfamily in Zebrafish. BMC Evol. Biol. 8, 42. doi:10.1186/1471-2148-8-42
Leeds, T. D., Silverstein, J. T., Weber, G. M., Vallejo, R. L., Palti, Y., Rexroad, C. E.,
et al. (2010). Response to Selection for Bacterial Cold Water Disease Resistance in
Rainbow Trout. J. Animal Sci. 88, 1936–1946. doi:10.2527/jas.2009-2538
Li, H. (2013). Aligning Sequence Reads, Clone Sequences and Assembly Contigs
with BWA-MEM. arXiv:1303.3997v2.
Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., et al. (2009).
The Sequence Alignment/Map Format and SAMtools. Bioinformatics 25,
2078–2079. doi:10.1093/bioinformatics/btp352
Liu, S., Vallejo, R. L., Palti, Y., Gao, G., Marancik, D. P., Hernandez, A. G., et al.
(2015). Identi fication of Single Nucleotide Polymorphism Markers Associated
with Bacterial Cold Water Disease Resistance and Spleen Size in Rainbow
Trout. Front. Genet. 6, 298. doi:10.3389/fgene.2015.00298
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 93680611
Liu et al. Haplotypes Associated With BCWD Resistance

Liu, S., Gao, G., Layer, R. M., Thorgaard, G. H., Wiens, G. D., Leeds, T. D., et al.
(2021). Identification of High-Confidence Structural Variants in Domesticated
Rainbow Trout Using Whole-Genome Sequencing. Front. Genet. 12, 639355.
doi:10.3389/fgene.2021.639355
Liu, S., Palti, Y., Gao, G., and Rexroad, C. E. (2016). Development and Validation of
a SNP Panel for Parentage Assignment in Rainbow Trout. Aquaculture 452,
178–182. doi:10.1016/j.aquaculture.2015.11.001
Liu, S., Palti, Y., Martin, K. E., Parsons, J. E., and Rexroad, C. E. (2017). Assessment
of Genetic Differentiation and Genetic Assignment of Commercial Rainbow
Trout Strains Using a SNP Panel. Aquaculture 468, 120–125. doi:10.1016/j.
aquaculture.2016.10.004
Liu,S.,Vallejo,R.L.,Evenhuis,J.P.,Martin,K.E.,Hamilton,A.,Gao,G.,etal.
(2018). Retrospective Evaluation of Marker-Assisted Selection for
Resistance to Bacterial Cold Water Di sease in Three Generations of a
Commercial Rainbow Trout Breeding Population. Fr ont. Genet. 9, 286.
doi:10.3389/fgene.2018.00286
Loch, T. P., and Faisal, M. (2015). Emerging Flavobacterial Infections in Fish: A
Review. J. Adv. Res. 6, 283–300. doi:10.1016/j.jare.2014.10.009
Marancik, D., Gao, G., Paneru, B., Ma, H., Hernandez, A . G., Salem, M., et al. (2015).
Whole-Body Transcriptome of Selectively Bred, Resistant-, Control-, and
Susceptible-Line Rainbow Trout Following Experimental Challenge with
Flavobacterium Psychrophilum. Front. Genet. 5, 453. doi:10.3389/fgene.2014.00453
Narum, S. R., Di Genova, A., Micheletti, S. J., and Maass, A. (2018). Genomic
Variation Underlying Complex Life-History Traits Revealed by Genome
Sequencing in Chinook Salmon. Proc. R. Soc. B-Biol. Sci. 285 (1883),
20180935. doi:10.1098/rspb.2018.0935
Naylor, R. L., Kishore, A., Sumaila, U. R., Issifu, I., Hunter, B. P., Belton, B., et al.
(2021). Blue Food Demand across Geographic and Temporal Scales. Nat.
Commun. 12, 5413. doi:10.1038/s41467-021-25516-4
Nematollahi, A., Decostere, A., Pasmans, F., and Haesebrouck, F. (2003).
Flavobacterium Psychrophilum Infections in Salmonid Fish. J. Fish. Dis. 26,
563–574. doi:10.1046/j.1365-2761.2003.00488.x
O’Connell, J. R., and Weeks, D. E. (1998). PedCheck: A Program for Identification
of Genotype Incompatibilities in Linkage Analysis. Am. J. Hum. Genet. 63,
259–266. doi:10.1086/301904
Palti,Y.,Gao,G.,Liu,S.,Kent,M.P.,Lien,S.,Miller,M.R.,etal.(2015a).The
Development and Characterization of a 57K Single Nucleotide Polymorphism Array
for Rainbow Trout. Mol. Ecol. Resour. 15, 662–672. doi:10.1111/1755-0998 .12337
Palti, Y., Vallejo, R. L., Gao, G., Liu, S., Hernandez, A. G., Rexroad, C. E., et al. (2015b).
Detection and Validation of QTL Affecting Bacterial Cold Water Disease
Resistance in Rainbow Trout Using Restriction-Site Associated DNA
Sequencing. PLoS One 10, e0138435. doi:10.1371/journal.pone.0138435
Pearse, D. E., Barson, N. J., Nome, T., Gao, G., Campbell, M. A., Abadía-Cardoso, A.,
et al. (2019). Sex-Dependent Dominance Maintains Migration Supergene in
Rainbow Trout. Nat. Ecol. Evol. 3, 1731–1742. doi:10.1038/s41559-019-1044-6
R Core Team (2021). R: A Language and Environment for Statistical Computing
[Online]. Vienna, Austria: R Foundation for Statistical Computing. Available:
https://www.R-project.org (Accessed Dec. 17, 2021).
Schlotterer, C., Tobler, R., Kofler, R., and Nolte, V. (2014). Sequencing Pools of
Individuals-Mining Genome-Wide Polymorphism Data without Big Funding.
Nat. Rev. Genet. 15, 749–763. doi:10.1038/nrg3803
Schröder, B. (2016). The Multifaceted Roles of the Invariant Chain CD74 - More
Than Just a Chaperone. Biochim. Biophys. Acta (BBA) - Mol. Cell Res.
1863,
1269–1281. doi:10.1016/j.bbamcr.2016.03.026
Semple, S. L., Heath, G., Christie, D., Braunstein, M., Kales, S. C., and Dixon, B.
(2019). Immune Stimulation of Rainbow Trout Reveals Divergent Regulation of
MH Class II-Associated Invariant Chain Isoforms. Immunogenetics 71,
407–420. doi:10.1007/s00251-019-01115-y
Song, Z., Zou, J., Wang, M., Chen, Z., and Wang, Q. (2022). A Comparative Review
of Pyroptosis in Mammals and Fish. J. Inflamm. Res. 15, 2323–2331. doi:10.
2147/jir.s361266
Starliper, C. E. (2011). Bacterial Coldwater Disease of Fishes Caused by Flavobacterium
Psychrophilum. J. Adv. Res. 2, 97–108. doi:10.1016/j.jare.2010.04.001
Therneau, T. M. (2021). A Package for Survival Analysis in R. R Package Version
3.2-13. [Online] Available: https://CRAN.R-project.org/package=survival
(Accessed Dec. 17, 2021).
Thompson,N.F.,Anderson,E.C.,Clemento,A.J.,Campbell,M.A.,Pearse,D.E.,
Hearsey, J. W., et al. (2020). A Complex Phenotype in Salmon Controlled by a
Simple Change in Migratory Timing. Science 370, 609–613. doi:10.1126/science.
aba9059
Vallejo, R. L., Leeds, T. D., Gao, G., Parsons, J. E., Martin, K. E., Evenhuis, J. P., et al.
(2017a). Genomic Selection Models Double the Accuracy of Predicted Breeding
Values for Bacterial Cold Water Disease Resistance Compared to a Traditional
Pedigree-Based Model in Rainbow Trout Aquaculture. Genet. Sel. Evol. 49, 17.
doi:10.1186/s12711-017-0293-6
Vallejo, R. L., Liu, S., Gao, G., Fragomeni, B. O., Hernandez, A. G., Leeds, T. D., et al.
(2017b). Similar Genetic Architecture with Shared and Unique Quantitative Trait
Loci for Bacterial Cold Water Disease Resistance in Two Rainbow Trout Breeding
Populations. Front. Genet. 8, 156. doi:10.3389/fgene.2017.00156
Vallejo, R. L., Cheng, H., Fragomeni, B. O., Gao, G., Silva, R. M. O., Martin, K. E., et al.
(2021). The Accuracy of Genomic Predictions for Bacterial Cold Water Disease
Resistance Remains Higher Than the Pedigree-Based Model One Generation after
Model Training in a Commercial Rainbow Trout Breeding Population.
Aquaculture 545, 737164. doi:10.1016/j.aquaculture.2021.737164
Vallejo, R. L., Palti, Y., Liu, S., Evenhuis, J. P., Gao, G., Rexroad, C. E., et al. (2014b).
Detection of QTL in Rainbow Trout Affectin g Survival When Challenged with
Flavobacterium Psychrophilum. Mar. Biotechnol. 16, 349–360. doi:10.1007/
s10126-013-9553-9
Vallejo, R. L., Palti, Y., Liu, S., Marancik, D. P., and Wiens, G. D. (2014a).
Validation of Linked QTL for Bacterial Cold Water Disease Resistance and
Spleen Size on Rainbow Trout Chromosome Omy19. Aquaculture 432,
139–143. doi:10.1016/j.aquaculture.2014.05.003
Vallejo, R. L., Silva, R. M. O., Evenhuis, J. P., Gao, G., Liu, S., Parsons, J. E., et al.
(2018). Accurate Genomic Predictions for BCWD Resistance in Rainbow Trout
Are Achieved Using Low-Density SNP Panels: Evidence that Long-range LD is a
Major Contributing Factor. J. Anim. Breed. Genet. 135, 263–274. doi:10.1111/jbg.
12335
Veillette, A. (2006). Immune Regulation by SLAM Family Receptors and SAP-
Related Adaptors. Nat. Rev. Immunol. 6, 56–66. doi:10.1038/nri1761
Wiens, G. D., Lapatra, S. E., Welch, T. J., Evenhuis, J. P., Rexroad, C. E., and Leeds,
T. D. (2013a). On-Farm Performance of Rainbow Trout (Oncorhynchus
Mykiss) Selectively Bred for Resistance to Bacterial Cold Water Disease:
Effect of Rearing Environment on Survival Phenotype. Aquaculture 388-391,
128–136. doi:10.1016/j.aquaculture.2013.01.018
Wiens,G.D.,Palti,Y.,andLeeds,T.D.(2018).ThreeGenerationsofSelectiveBreeding
Improved Rainbow Trout (Oncorhynchus Mykiss) Disease Resistanc e against
Natural Challenge with Flavobacterium Psychrophilu m during Early Life-Stage
Rearing. Aquaculture 497, 414–421. doi:10.1016/j.aquaculture.2018.07.064
Wiens, G. D., Vallejo, R. L., Leeds, T. D., Palti, Y., Hadidi, S., Liu, S., et al. (2013b).
Assessment o f Genetic Correlation betwee n Bacterial Cold Water Disease Resistance
and Spleen Inde x in a Domesticated Population of Rainbow Trout: Identification of
QTL on Chromosome Omy19. PLoS One 8, e75749. doi:10.1371/journal.pone.0075749
Xiang, R. D., Macleod, I. M., Daetwyler, H. D., De Jong, G., O’connor, E.,
Schrooten, C., et al. (2021). Genome-Wide Fine-Mapping Identifies
Pleiotropic and Functional Variants that Predict Many Traits across Global
Cattle Populations. N at. Commun. 12, 860. doi:10.1038/s41467-021-21001 -0
Conflict of Interest: KM was employed by the company Troutlodge Inc.
The remaining authors declare that the research was conducted in the absence of
any commercial or financial relationships that could be construed as a potential
conflict of interest.
Publisher’s Note: All claims expressed in this article are solely those of the authors
and do not necessarily represent those of their affiliated organizations, or those of
the publisher, the editors and the reviewers. Any product that may be evaluated in
this article, or claim that may be made by its manufacturer, is not guaranteed or
endorsed by the publisher.
Copyright © 2022 Liu, Martin, Gao, Long, Evenhuis, Leeds, Wiens and Palti. This is
an open-access article distributed under the terms of the Creative Commons
Attribution License (CC BY). The use, distribution or reproduction in other
forums i s perm itted, provided the origin al au thor(s) and the copyright owner(s)
are credited and that the original publication in this journal is cited, in
accordance with accepted academic practice.Nouse,distributionor
reproduction is permitted whi ch does not comply with these terms.
Frontiers in Genetics | www.frontiersin.org June 2022 | Volume 13 | Article 93680612
Liu et al. Haplotypes Associated With BCWD Resistance
